Measurements and Scalar Data for Multi-ROIs
You can compute measurements and add other scalar data to multi-ROIs in the Compute Measurements dialog and from the multi-ROI contextual menu. You can also import, map, and copy scalar values to multi-ROIs, as well as export scalar values from multi-ROIs.
Compute Measurements dialog
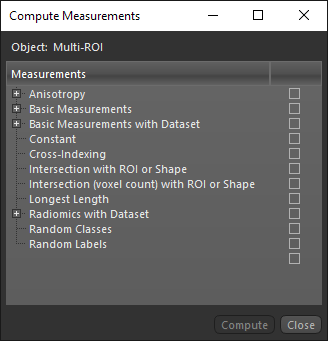
The following measurements and other scalar values are available for multi-ROIs in the Compute Measurements dialog.
| Description | |
|---|---|
| 2D Measurements |
Lets you compute measurements, such as the area, filled area, perimeter, solidity, and compactness, of the discrete objects within a 2D multi-ROI.
Note Refer to the topic 2D Measurements for descriptions of the available measurements. |
| Anisotropy |
Lets you compute anisotropy for the classes within a multi-ROI using the mean intercept length (MIL) or star volume distribution (SVD) method.
Note Refer to the topic for Anisotropy for descriptions of the available methods and settings for computing anisotropy. |
| Basic Measurements |
Lets you compute measurements, such as surface areas, aspect ratios, centers of mass, and other parameters, of the discrete objects within a multi-ROI.
Note Refer to the topic Basic Measurements for descriptions of the available measurements. |
| Basic Measurements with Dataset |
Lets you compute values, such as minimum, mean, maximum intensity values, weighted centers of mass, and entropy, corresponding to the selected image data and the labeled voxels of the discrete objects in a multi-ROI.
Note Refer to the topic Basic Measurements with Dataset for descriptions of the available measurements. |
| Constant |
Lets you add a constant value to each class within a multi-ROI. You can choose a measurement title and constant value, as shown below.
|
| Cross-Indexing |
Lets you cross-index the label number of a multi-ROI's classes to the intersecting classes of a selected multi-ROI. As shown below, you can choose a measurement title and multi-ROI to cross-index.
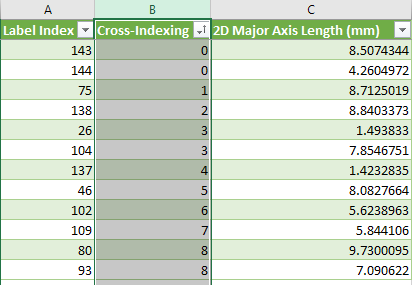
As shown in the example below, the Cross-Indexing column shows the intersecting label index of the selected multi-ROI. You should note that a value of 0 indicates that the label of the selected multi-ROI does not intersect with any labels of the multi-ROI. In addition, in cases in which a label intersects with multiple labels, the first intersecting label will be reported.
|
|
Intersection with ROI |
Lets you identify classes that intersect with a selected region of interest
The result is either 1 or 0 — intersecting or not. |
|
Intersection (voxel count) with ROI |
Computes the number of pixels within a multi-ROI's classes that intersect with a selected region of interest |
| Longest Length | Two computations are available for Longest Length — Longest Length and Tortuosity. These parameters are often used as part of the analysis of fibered datasets.
Longest Length… Computes the longest length of each labeled object in the multi-ROI. For example, as part of the analysis of a fibers dataset. Tortuosity… Computes the tortuosity of each labeled object in the multi-ROI. |
| Radiomics with Dataset | Lets you extract quantitative features from images using data-characterization algorithms. Once features are extracted, computational algorithms and statistical models can be applied to analyze and interpret the extracted data.
The various features that can be extracted using radiomics, on either an original or derived image, can be subdivided into the following classes: Gray Level Co-occurrence Matrix (GLCM)… These 24 features provide for quantitative and objective analyzes of image textures (go to https://pyradiomics.readthedocs.io/en/latest/features.html#module-radiomics.glcm for more information about GLCM). Gray Level Dependence Matrix (GLDM)… The 14 features of the GLDM provide information about the coarser-scale texture properties of an image compared to GLCM features. The strength and direction of gray-level dependencies captured and can be particularly useful for analyzing patterns characterized by variations in gray-level values (go to https://pyradiomics.readthedocs.io/en/latest/features.html#module-radiomics.gldm for more information about GLDM). Gray Level Run Length Matrix (GLRLM)… These 16 features capture information about the spatial arrangement and distribution of pixel runs in an image. Providing insights into the coarseness, regularity, and complexity of underlying textures, these features can identify regions with specific texture patterns for image segmentation (go to https://pyradiomics.readthedocs.io/en/latest/features.html#module-radiomics.glrlm for more information about GLRLM). Gray Level Size Zone Matrix (GLSZM)… The 16 features of the GLSZM provide information about the distribution and characteristics of connected regions within an image by capturing the heterogeneity, fragmentation, and spatial organization of texture patterns. Theses features can identify regions of interest with specific size and texture characteristics for image segmentation (go to https://pyradiomics.readthedocs.io/en/latest/features.html#module-radiomics.glszm for more information about GSZM). Neighboring Gray Tone Difference Matrix (NGTDM)… The five features of the NGTDM provide information about the local texture variations and gradients in an image by capturing the differences in gray-level values between adjacent pixels to reveal fine-scale details of texture patterns. They can also identify regions of interest with specific local texture variations for image segmentation (go to https://pyradiomics.readthedocs.io/en/latest/features.html#module-radiomics.ngtdm for more information about NGTDM). |
| Random Classes |
Adds random classes to a multi-ROI. You can enter a title for the values and choose the number of classes, as shown below.
|
| Random Labels | Adds random labels to a multi-ROI. You can enter a title for the values and the number of classes required. |